Python in Science: How long until a Nobel Prize?
As I write this, the Nobel Prizes for 2007 are being announced. During the week of announcements, each day includes news of another award being bestowed for outstanding contributions in physics, chemistry, physiology or medicine, literature, peace, and economics. As a technophile, the science awards have always been the most interesting to me. This year, prior to the awards, new releases of several scientific packages on PyPI caught my eye and I was struck by the coincidence. I started to wonder: How long before a Nobel Prize is awarded to a scientist who uses Python for their work in some significant way?
It should come as no surprise that Python can be found in many scientific environments. The language is powerful enough to do the necessary work, and simple enough to allow the focus to remain on the science, instead of the programming. It is portable, making sharing code between researchers easy. Its ability to interface with other systems, through C libraries or network protocols, also makes Python well suited for building on existing legacy tools. The list of tools highlighted here is by no means exhaustive, but is intended to introduce the wide variety of packages available for scientific work in Python, ranging from general mathematics tools to narrowly focused, specialty libraries.
The home for scientific programming in Python is the SciPy home page. SciPy aims to collect references to all of the scientific libraries and serve as a central hub for sharing code in the scientific programming community. There is even a SciPy conference! The SciPy site is an excellent source of information about scientific programming in general, and using Python specifically. It’s a good starting point for a survey to examine how Python is used in scientific research work.
General Purpose Toolkits
Most scientific work includes some data collection and management, along with number crunching to analyze that data. There are several general purpose libraries for working with datasets from any scientific field.
The NumPy library is designed as “the fundamental package needed for scientific computing with Python”. It includes powerful data management and manipulation features, including multi-dimensional arrays. Once your data is collected, it can be processed by NumPy using linear algebra, Fourier transforms, a random number generator, and FORTRAN integration. NumPy is the foundation of several libraries also hosted on or otherwise associated with the SciPy site.
The main SciPy library uses the array manipulation features of NumPy to implement more advanced mathematical, scientific, and engineering functions. It provides routines for numerical integration and optimization, signal processing, and statistics.
PyTables is a data management package specifically designed to handle large datasets. It manages data file access with the HDF5 library, and uses NumPy for in-memory datasets. Since it is designed for use with large amounts of data, it is well suited for applications which produce or collect a lot of data, even outside the scientific arena.
The ScientificPython package is another broad ranging collection of modules for scientific computing. It includes an input/output library; geometric, mathematical, and statistical functions; as well as several general purpose modules to assist in programming tasks such as threading and parallel programming.
Simulations
Besides working with observed data, scientists often use simulations to understand the rules governing the operation of a system. If you can construct a simulation that accurately predicts the outcome of a set of inputs, then you have a higher level of confidence that you understand the way the parts of a system interact. There are basically two types of simulation, discrete and continuous, and there are Python packages for working with both.
SimPy is a simulation language based on standard Python. You can use SimPy to simulate activities like traffic flow patterns, queues at retail stores, and other discrete events. SimPy represents independent active components of the simulation as “processes” that can interact with each other (queuing, passing data or resources, etc.). Limited capacity is represented through “resources”, and requests for resources are maintained for you.
PyDSTool is a suite of computational tools for modeling dynamic systems and physical processes being developed at Cornell. It supports discrete or continuous simulation, and a wide range of mathematical operations and constraints. One especially interesting feature is the use of automatically generated and compiled C code for the “generators” that produce input data used by the rest of the tool.
Mathematics
Once your data is collected, it needs to be analyzed. That may involve statistical calculations, or you might be trying to uncover an underlying relationship or formula. In either case, there is a Python library with the tools you need.
SymPy – not to be confused with SimPy – is a full featured computer algebra system, for symbolic mathematics. It supports algebraic formula expansion and reduction, as well as calculus operations such as differentials, derivatives, series and limits, and integration.
If you need even more powerful mathematics, or just want to take advantage of previous work done with the ubiquitous MATLAB, you can use pymatl to drive the MATLIB engine directly from your Python program. It will send matrices of data back and forth between your Python program and MATLIB, allowing the two programs to act together on the data.
Visualization
Once your data is collected and you have completed your calculations, the next step is generally to produce graphical views of the data to make it easier to interpret the results and spot trends. The 2-D plotting and visualization matplotlib is used in a lot of different application areas. It produces publication quality figures in a variety formats, and you can control it from scripts, GUIs, through the web, or interactively through the python or ipython shells. Figure 1 shows a sample plot created with matplotlib by Jeff Whitaker of NOAA’s Earth System Research Lab in Boulder, CO (and author of the basemap toolkit for matplotlib).
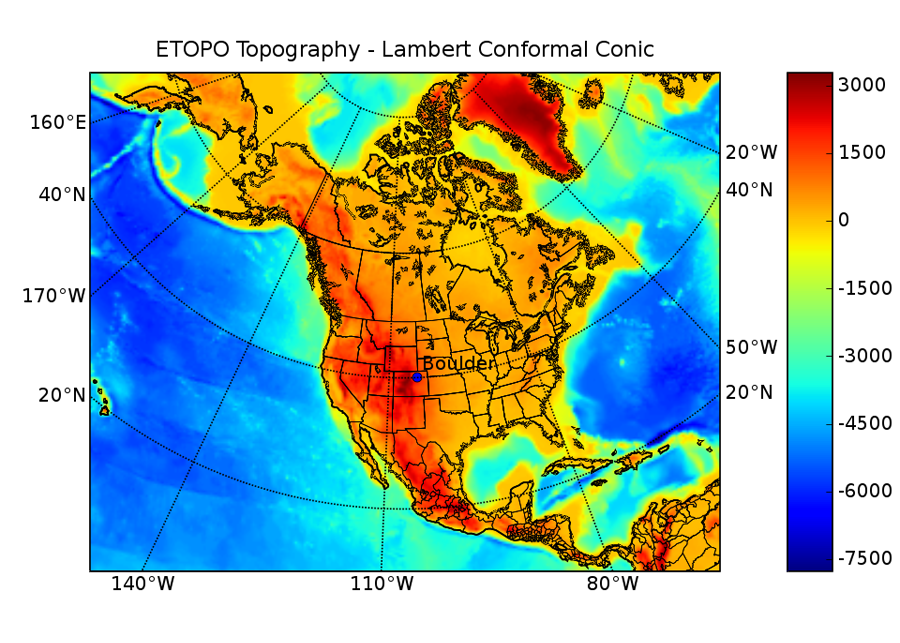
A matplotlib example from Jeff Whitaker.
For 3-D visualization, you will want to check out Mayavi, from Enthought. Mayavi is a general purpose visualization engine, and it can be used with scalar or vector data to create and manipulate three dimensional representations of your dataset for visualization.
Astronomy
In addition to the many general purpose libraries suitable for scientific work, there are quite a few application specific libraries available in different fields from the macroscopic to the sub-microscopic.
The AstroPy project from the Astronomy Department at the University of Washington in Seattle promotes the use of Python for astronomy research. Their home page lists several packages for accessing legacy systems through Python wrappers, as well as pure Python libraries for working with astronomical data. For example, AstroLib is a collection of 4 components which provide features such as manipulating the ASCII tables commonly used to exchange data between scientists, synthetic photometry (for analyzing intensity or apparent magnitude measurements), and coordinate conversion and manipulation. The target user is a “typical astronomer” preparing for observation runs or working with observed or catalog data.
PyNOVAS is a library for calculating the positions of the sun, moon, planets, and other celestial objects. It is based on a C library, called NOVAS, and is a good example of wrapping existing libraries in Python to make them available to a wider range of scientists.
The Space Telescope Science Institute manages the operation of the Hubble Space Telescope. In addition to using Python for many of their internal tools, they have released a library for working with astronomical data and images, called stsci_python.
Climate
If astronomy isn’t your thing, maybe you want to look at scientific applications a little closer to home. Climate research is a hot topic these days, both in the news and in Python development.
The tools in the Climate Data Analysis Tools (CDAT) system from Lawrence Livermore National Laboratory are specifically designed for working with climate data. There are separate components for reading and writing data, performing climate-specific calculations, and general statistical analysis.
PyClimate from Universidad del País Vasco in Spain is focused on analyzing and modeling climate data, combining data from different sources in different formats and measurements to look for variability, especially human induced change.
Bjørn Ådlandsvik’s seawater module implements functions for computing properties of the ocean, using standard formulas defined by UNESCO reports, while the fluid package is a more general set of procedures for studying fluid interactions.
Biology / Health
If you are more interested in animated creatures than inanimate objects, you will be pleased to know that there is a thriving community of biology researchers using Python for their work.
Biopython hosts a set of tools for “computational molecular biology” for bioinformatics. Contributors are distributed around the world, mostly at research universities. The library they have produced includes tools for parsing bioinformatics files from a wide range of sources, offline and online. It also contains classes for representing and manipulating DNA sequences.
EpiGrass is used for network epidemiology simulations. The results can be fed to the GRASS geographic information system and plotted on maps to track or predict the spread of disease.
Molecular Modeling
If we continue this trend toward studying smaller objects, we soon reach the microscopic and sub-microscopic scales and find Python hard at work there, too.
The Scripps Research Institute has released several tools for visualizing and analyzing molecular structures. Their Python Molecular Viewer draws an interactive three dimensional representation of a molecule. It is also scriptable, using built-in or user defined commands dynamically loaded from plug-ins.
The Molecular Modeling Toolkit by Konrad Hinsen is another simulation application, this time specifically intended for simulating molecules and their interactions.
Chimera, from the University of California, San Francisco, is an alternate interactive visualization and analysis tool for molecular structures. It produces high quality images and animations, and can be driven by a command interface or interactively.
Conclusion
Although I have barely scratched the surface, I hope this list of packages illustrates the wide array of application areas where Python is being used for scientific research. From the macroscopic to microscopic, simulation to computation, it fills gaps left by other tools and serves as the foundation technology for entirely new tools. Whether it is reading and writing standard (or ad hoc) data files, controlling equipment, or performing calculations directly, Python has an important place in science. It can take several decades before the impact of fundamental research is evaluated and recognized as worthy of a Nobel Prize, and Python is still young enough that the research being done using needs to mature before it would be considered. But that day is coming, and it is entirely possible that a Nobel Prize will be awarded to a scientist who uses Python within my lifetime.
Update on the GIL
Thanks to everyone who sent a message or link after last month’s column! The responses were generally positive. One of the corrections came from Adam Olsen, who reported that he is working on a branch of the C interpreter which removes the GIL. I missed the discussion on the python-dev list, so I wasn’t aware of his project. According to Adam, the code is pre-alpha status, and still needs work in areas such as deadlock management and weakrefs. He does have some performance numbers, gathered using the pystone benchmarks running on a dual core system. As a baseline, an unmodified version of Python 3000 yields 28000 pystones/second. His GIL-free version produces 18800 pystones/second for one thread, and 36700 pystones/second for two threads.
As always, if there is something you would like for me to cover in
this column, send a note with the details to doug dot hellmann at
pythonmagazine dot com and let me know, or add the link to your
del.icio.us account with the tag pymagdifferent.